Select analysis version to view the applicable content:
Now that we have talked about meaningful statistics in impedance cardiography and how they are calculated, we need to take a step back and make sure the data are clean and free of artifact. There are two steps for editing in impedance cardiography – R peak editing and Ensemble editing.
R Peak Editing
Recall from last time that many of these statistics are derived from the Ensemble Average, which is an average depiction of the cardiac cycle dependent on the detection of the R peak on the ECG waveform. If the R peaks aren’t detected properly, the Ensemble Average won’t be built correctly and the statistics will be incorrect.
I won’t cover ECG editing techniques much here, as this has been discussed in depth in previous posts. What is important to note from those posts is that for Impedance Cardiography, only mark normal R peaks with good corresponding dZ/dt cycles. This is different than editing ECG for Heart Rate Variability, where a contiguous R peak series is needed.
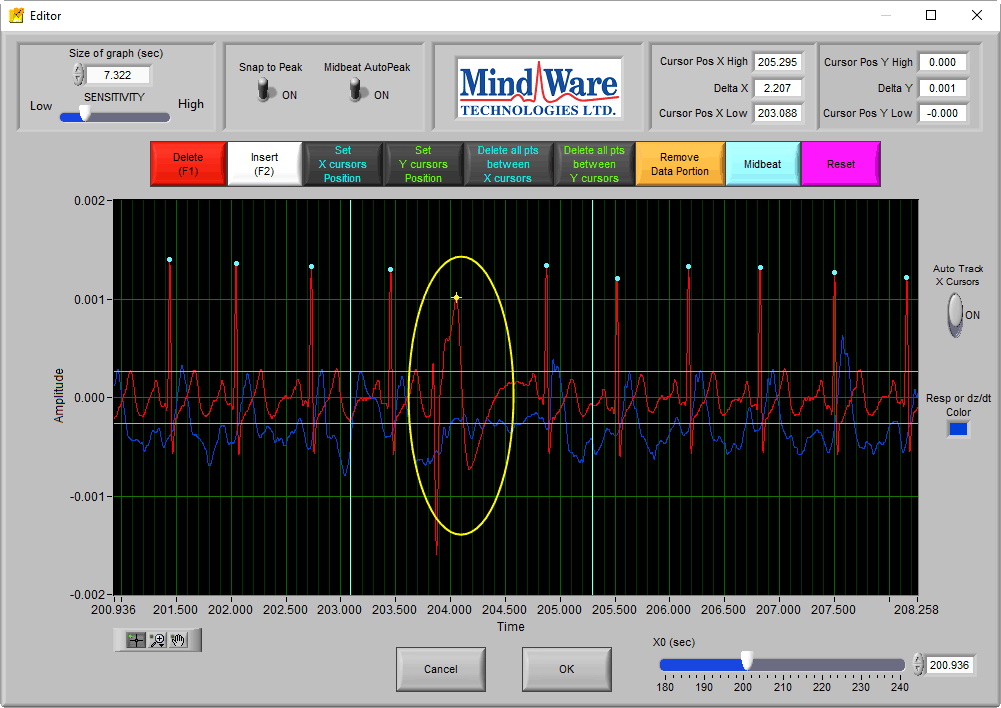
Take for example the following abnormal beat:
If this peak was left marked, it would be used to build the Ensemble Average. But it is clearly not actually an R peak, so the first thing to do would be to delete this peak.
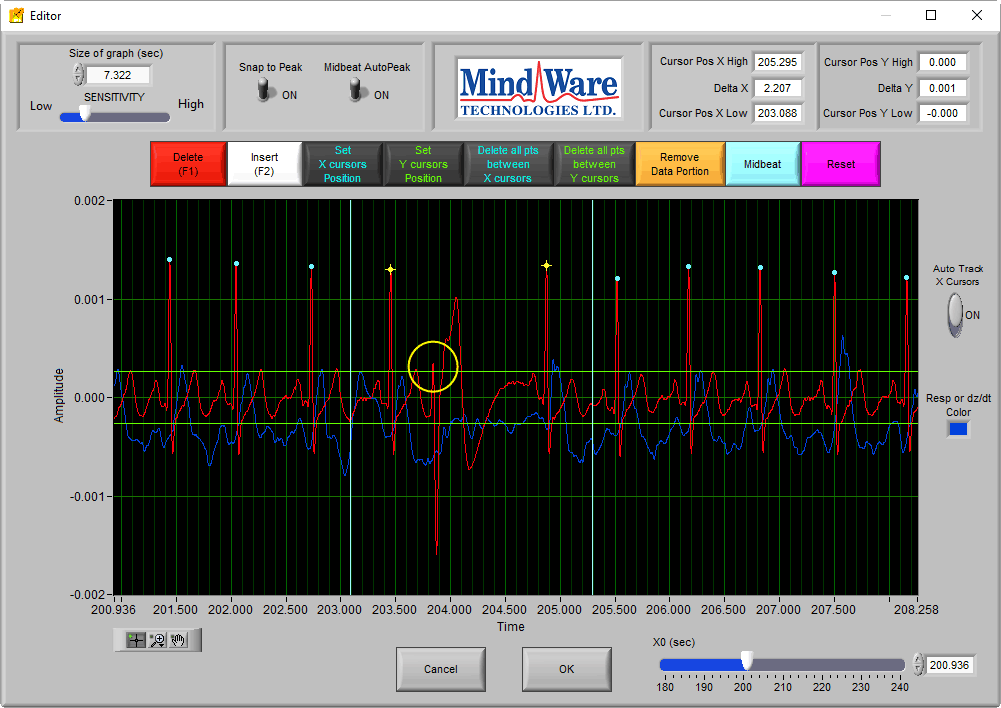
Now we can see something that looks like an abnormal R peak. If we were to mark this peak, it would corrupt the Ensemble Average by including an ECG cycle that does not match the expected signal morphology for the segment. Additionally, there is no dZ/dt response following the pattern of previous beats. In this case, no peak should be inserted.
If this data were edited for HRV, you may see an Absolute Midbeat placed between the two normal beats to maintain timing.
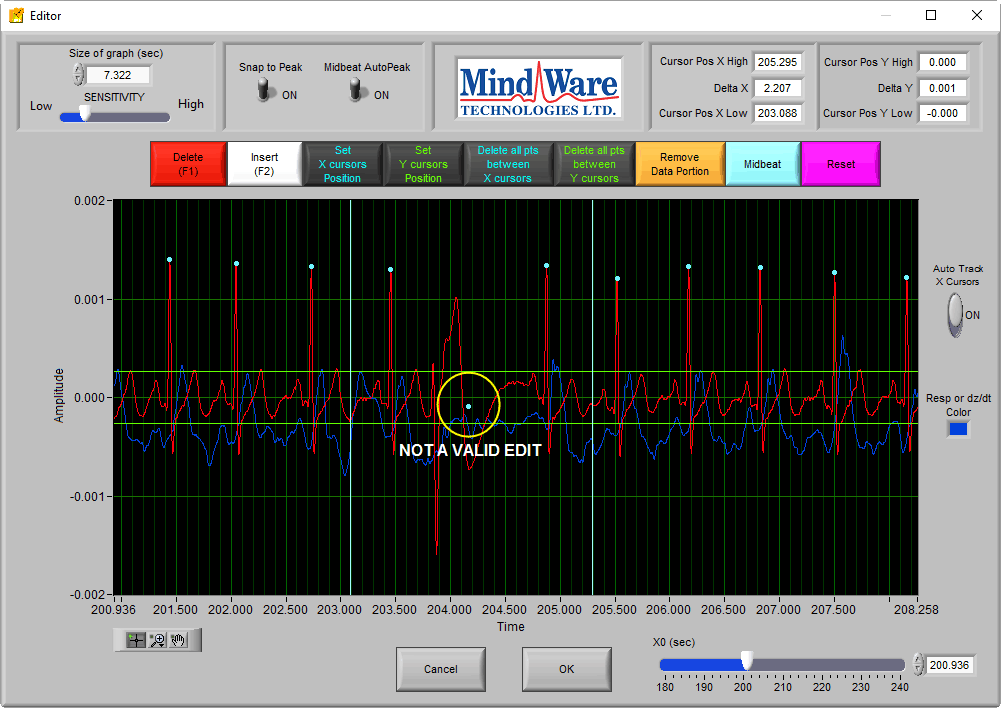
This type of edit is not valid for impedance cardiography, for the same reasons as above – corruption of the Ensemble Average by including data that does not depict a normal cardiac cycle.
Take for example the following abnormal beat:
If we left this peak marked, it would corrupt the Ensemble Average by including an ECG cycle that does not match the expected signal morphology for the segment. Additionally, there is no dZ/dt response following the pattern of previous beats. In this case, we should delete this peak.
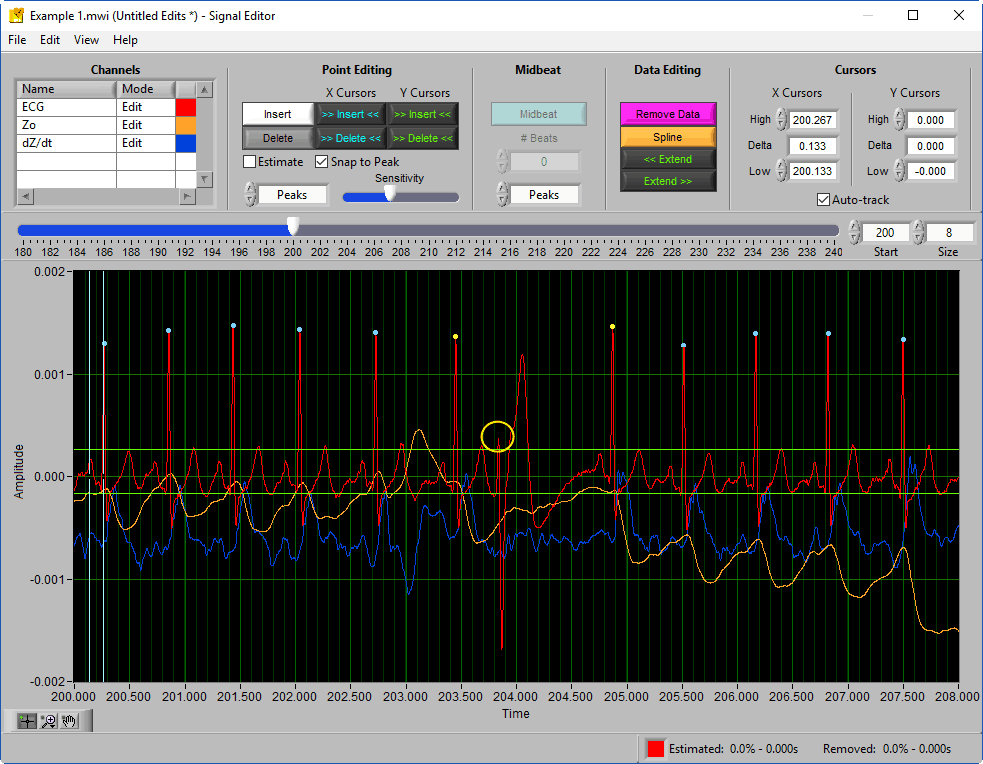
If this data were edited for HRV, you may see an Absolute Midbeat placed between the two normal beats to maintain timing.
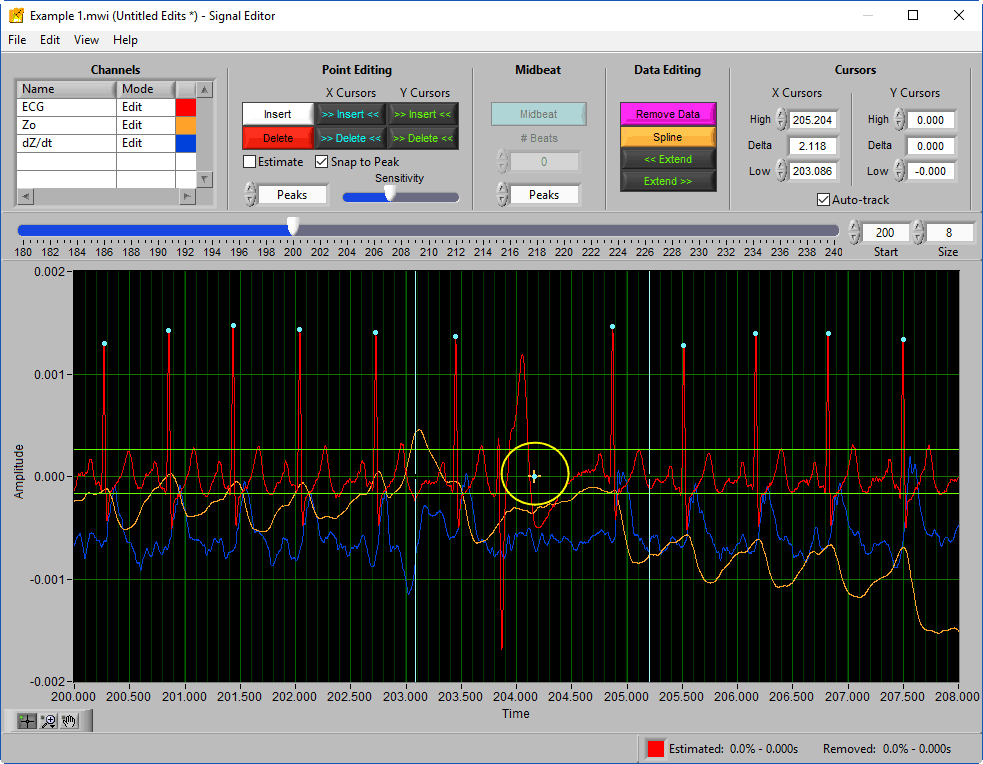
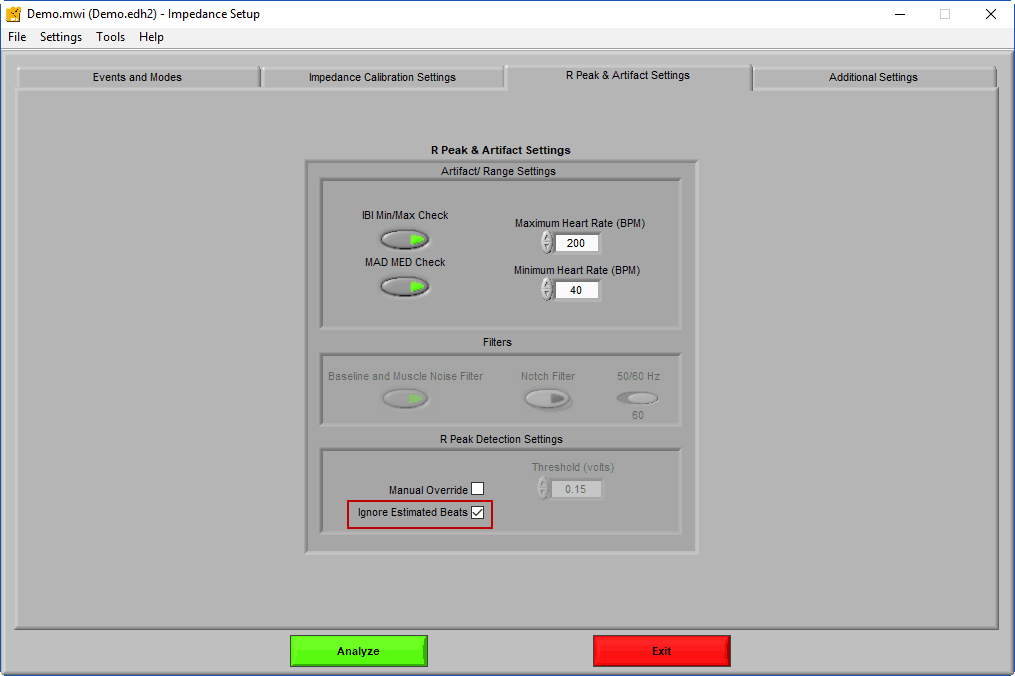
This is not a valid edit for impedance analysis, but if you are re-using edits from HRV (or planning on using these edits in HRV) it is OK to leave this estimated peak as long as Ignore Estimated Beats was enabled on the Setup screen.
Ensemble Editing
Once the ECG signal has been edited and we know the data being used to build the Ensemble Average is correct, it is time to examine the Ensemble Average itself. The algorithms used to detect landmarks on the ECG and dZ/dt signals are not perfect – this is especially the case when the known characteristics for a given landmark are not present in the signal morphology due to noise or recording technique.
In the data file depicted below, the B point notch is visible in all segments so the Max Slope Change method for B Point Calculation was used. In this particular segment, the B point was not identified properly:
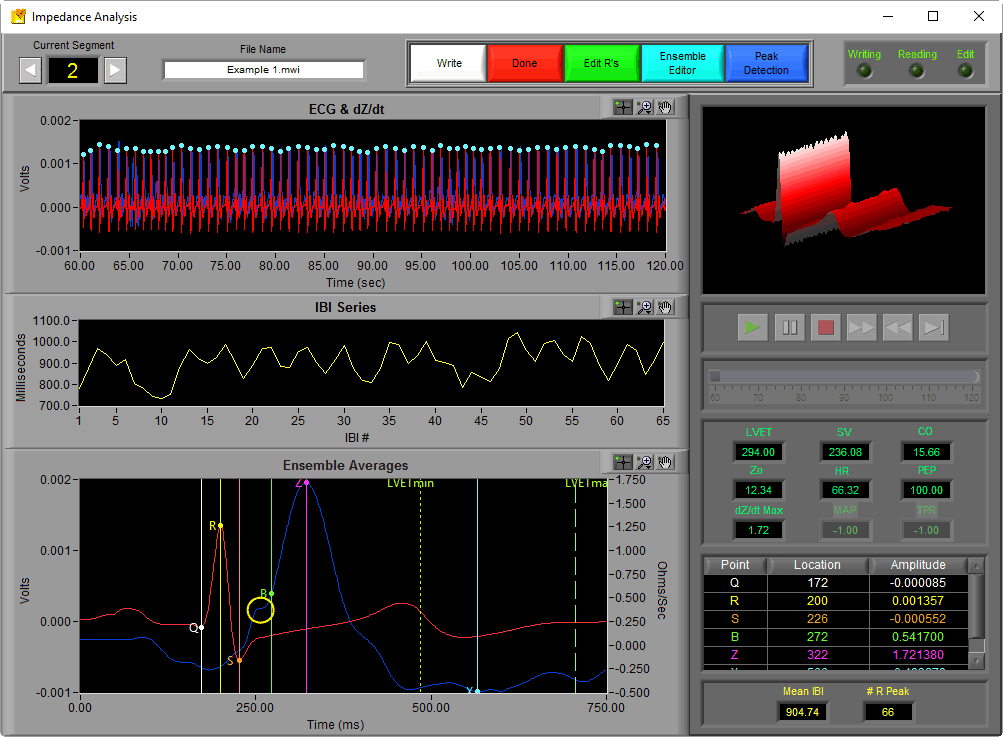
Since the B point was not placed properly in this instance, we should correct the B point placement using the Ensemble Editor.
Ensemble Editor
The Ensemble Editor allows you to move the points on the Ensemble Average to their correct locations in the event that they aren’t placed there automatically by the calculation algorithms. To launch the Ensemble Editor, press the light blue button near the top of the screen
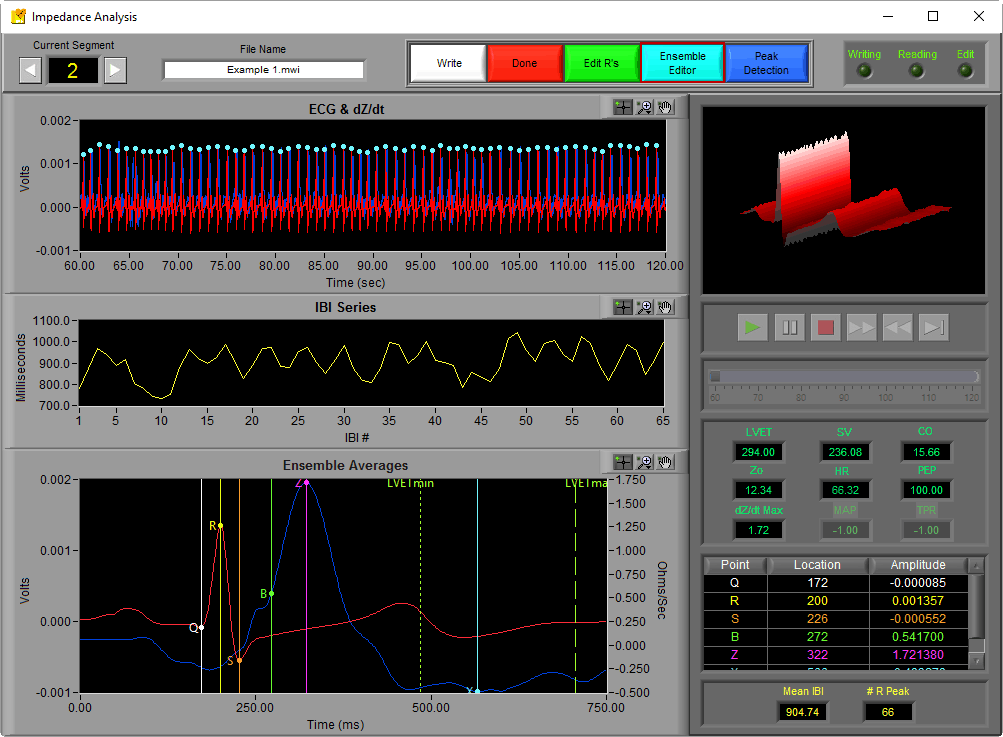
Once launched, the editor contains the Ensemble Average along with the location and amplitude of each of the landmarks
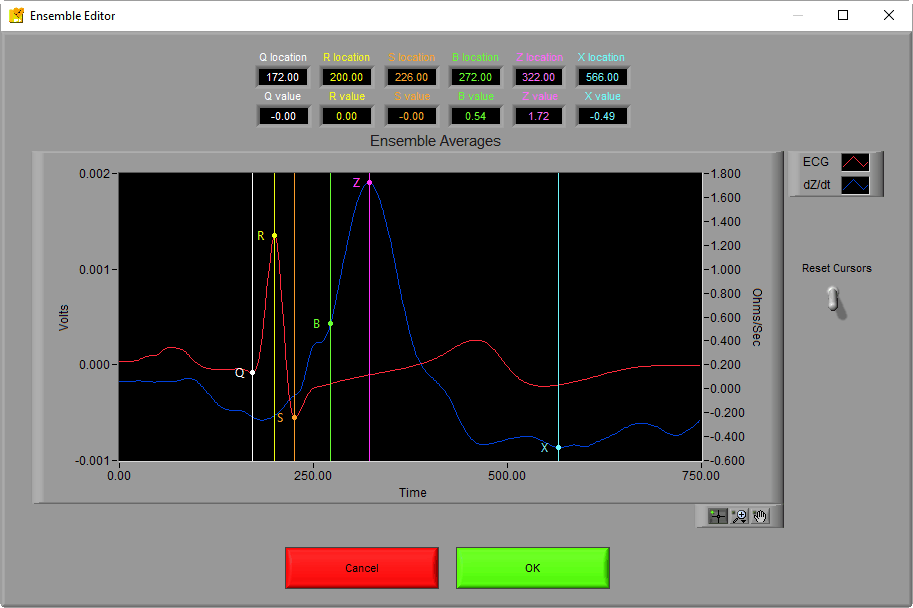
Ensure that the Crosshair tool is selected and click and drag any of the vertical lines to move the points to the desired locations. In this case, we want to drag the green B point to the left so that it rests in the notch
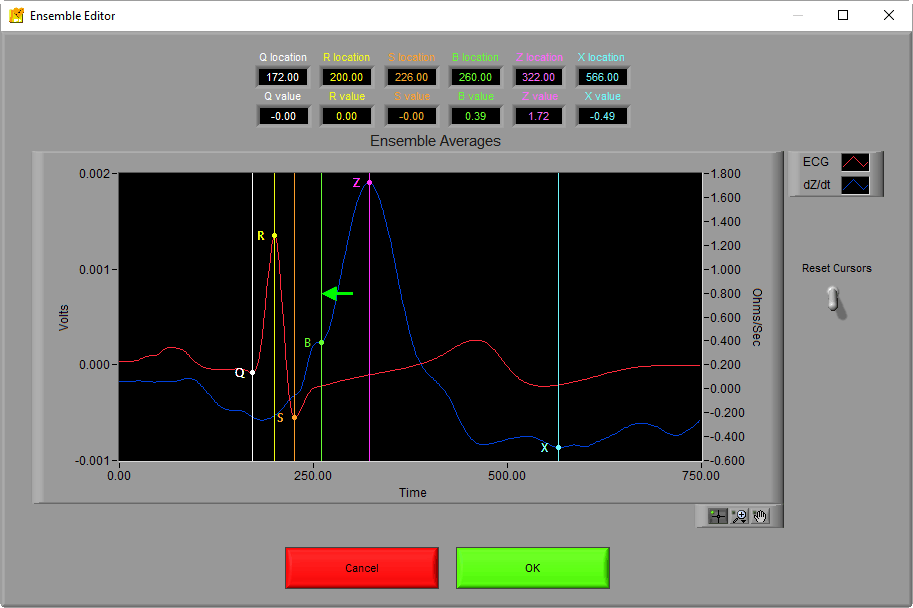
This can be done for any of the points that you feel have not been placed properly. If you need to start over, you can either reset the cursors using the lever to the right of the graph or press Cancel.
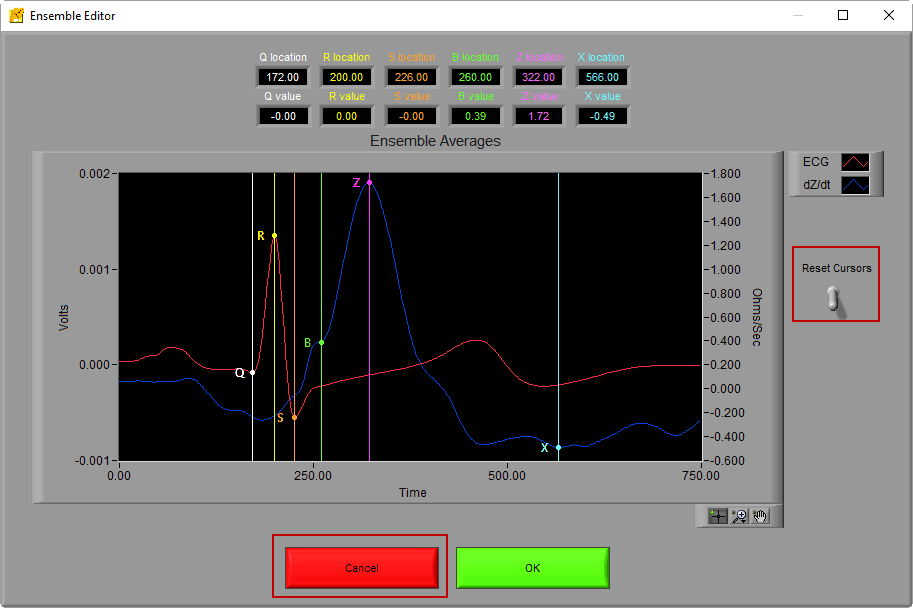
If you are satisfied with the changes, press OK to return to the IMP Analysis screen.
Adjust Point Identification Algorithm or Edit Point Placement?
In some situations, it makes more sense to adjust the algorithm used to place the landmark than editing the landmark itself. I am going to use the B point as an example for when it makes sense to adjust one versus the other.
Case 1
If you are working with a data set where the B points are visible, you should be using the Max Slope or Max Slope Change B Point Calculation methods. In this situation, if a B point is not placed exactly right, you should use the Ensemble Editor to adjust its location.
Case 2
If your data set does not have a clear B point visible, you have no choice but to estimate using Percent dZ/dt or Percent dZ/dt + C. Since there is no visible B point, you would never adjust the B point location.
Case 3
If your data set is somewhere in the middle, where some segments show the B point and some do not, it is actually best to estimate your B point location using one of the recommended methods. You can use the segments where B is visible to better inform what constant offset is used in the Percent dZ/dt + C method. Take the screenshot from above, and assume that B is being estimated

You can use the Ensemble editor to determine the offset between the estimated location and the actual location. Using this known discrepancy, you can raise/lower your constant offset or adjust the percentage in the B Point Calculation method to better estimate the B point location on segments where it isn’t visible.
Edit Files
There are two edit files stored by the IMP application – .edi and _ensemble.edt.
When selecting an edit file to use you will always choose the .edi file. If there are ensemble edits they will automatically be loaded from the corresponding _ensemble.edt file.
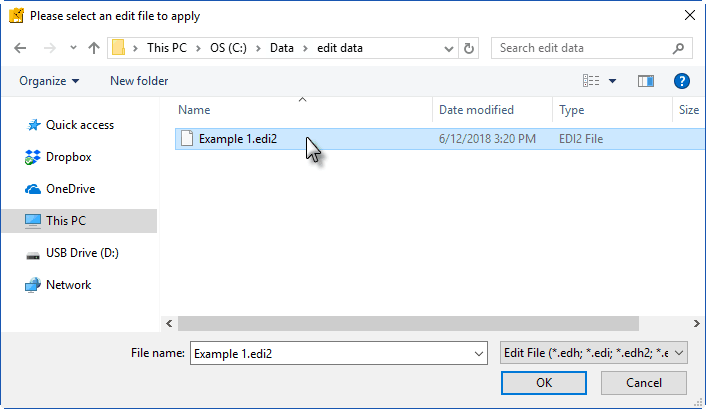
The edit file stored by the IMP application has a .edi2 file extension. This is the file you will select when specifying an edit file to use.
Helpful Hints
- Cleaning your ECG data is the foundation for a quality Ensemble Average and accurate statistics
- If re-using edits from HRV Analysis, be sure to watch out for estimated beats which should not be included
- Single beats have little influence on the Ensemble Average so, when in doubt as to whether an R peak is representative of a normal cardiac cycle, err on the side of deleting it
- Some forms of arrhythmia produce a decent ECG complex but no corresponding dZ/dt – these cases should also be discarded
- The algorithms are widely accepted and used, but they are not perfect. Use known visual cues to determine if the key points have been correctly placed
- If visual cues for a given landmark are not consistently visible, it is best to estimate. Use instances when visual evidence does exist to better inform your estimate
Next time we will wrap up our discussion on cardiac impedance by discussing how the signals can be used to derive respiration.